Building a ‘criminal database’ of environmental cancer causes
Jill Kucab from King’s College London explains how the Mutographs team are building a ‘criminal database’ of suspects that might be causing cancer.
Clues in our DNA
Sometimes people develop cancer and we have a pretty good idea what has caused it. For example, too much sun exposure can lead to skin cancer, and lung cancer is much more likely to develop in someone who smokes cigarettes. But can we actually prove it?
There are pieces of evidence hidden in the DNA code of tumour cells. These give us clues as to what might have caused a previously normal cell to change into cancerous one. Tumour cells contain many changes in their DNA code. These changes are called mutations. DNA is written using 4 letters (C, G, T and A) and a mutation can be as simple as one letter changing to another, like C becoming T. Some of these mutations are caused by chemicals or radiation in the environment.
For example, the DNA of skin cancers is full of C to T mutations. This change creates a ‘fingerprint’ of sun damage, also known as a signature. Smokers’ lung cancers have a large number of G to T mutations – more than we see in lung cancer from non-smokers. This G to T change is the ‘fingerprint’ caused by exposure to cigarette smoke.

Left: Fernando Caipa, Right: Pexels.com
Collecting evidence
But many times people are diagnosed with cancer and we don’t actually know what caused it. Is there a way to figure it out?
Our fellow scientists in Mutographs are sequencing the DNA from tumours donated by people with cancer around the world and studying the patterns of mutations that they contain. They use this information to try and discover what has caused these patterns. They think about what people may have been exposed to because of their lifestyle, where they live or where they work. Examples of the work that is being done include understanding why oesophageal cancer is more common in certain areas of the world, or what might cause the increased cases of liver cancer in Southeast Asia.
It is our hope that the DNA from these tumour types will contain clues that help us to figure out the cause. There may be fingerprints of environmental exposures, if we can find them. But just like uncovering fingerprints at a crime scene, we then need to figure out who (or actually what) the fingerprint belongs to. And to do that we need a reference set of fingerprints with known causes, like a criminal database for things that cause cancer.
Creating our ‘criminal database’
A key aspect of our work in the Environmental Carcinogenesis Group at King’s College London involves using cells grown in the lab to identify the fingerprints of things that are known or are suspected to cause cancer. We treat a number of different cell types with chemicals and radiation and study the DNA for patterns of mutations.

Picture credit: Halh Al-Serori
Sometimes we look at a single gene, which is a relatively short sequence of DNA (hundreds or thousands of letters), and sometimes we look at a person’s entire sequence of DNA (3 billion letters!). Looking at the all of the DNA in our cells helps us to detect a more complicated fingerprint.
Some experiments are performed using cells such as stem cells or mouse embryo fibroblasts. Both of these cell types grow well in the lab, making them useful for studying what happens to their DNA after different treatments. Using these cells we have confirmed that the changes to DNA detected in skin cancers are the same as what we see in cells exposed to ultraviolet radiation in the lab. We also showed that certain chemicals in cigarette smoke, such as benzo[a]pyrene, cause a mutational fingerprint in cells that is very similar to the fingerprint detected in smokers’ lung tumours. These data make us more confident that ultraviolet light is linked to skin cancer and that chemicals in cigarette smoke are linked to lung cancer. In addition to those examples, we have also identified fingerprints for many other chemicals using stem cells, including some found in diesel exhaust or the diet.
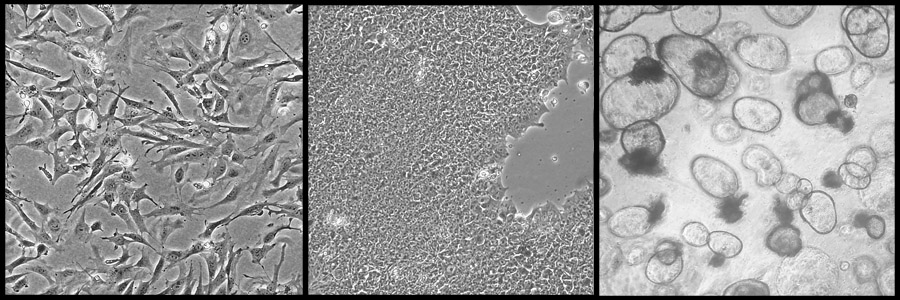
L-R: mouse embryo fibroblasts, stem cells, stomach organoids
Photo credit: Jill Kucab
More recently we have begun using organoids, or mini-organs. These are developed from donated tissues, such as the stomach, the intestine or the liver. While mouse embryo fibroblasts and stem cells are grown on plastic dishes as a single layer of cells, the organoids are grown as three-dimensional cultures and have a more complex structure. Mini-organs contain many of the different cell types that are found in the full sized-organ. They may also respond to chemicals in a way that better represents how the original tissue would respond. We are currently testing organoids developed from different tissues in the body to measure how they respond to treatment with potential cancer-causing agents and we are analysing their DNA to look for distinct patterns of changes. We can then compare the DNA from treated mini-organs with samples taken from people with cancer and see if they have similar ‘fingerprints’. These fingerprints might confirm exposure to a factor that was already suspected or could provide clues to exposures that were previously unknown.
Armed with this knowledge, we can then move onto the next stage which will be to work out how to prevent exposure to these factors and help people to reduce their risk of certain forms of cancer.